Note
Click here to download the full example code
06. 2D Interpolation¶
In the mathematical field of numerical analysis, interpolation is the problem of constructing new data points within the range of a discrete set of known data points. In signal and image processing, the data may be recorded at irregular locations and it is often required to regularize the data into a regular grid.
In this tutorial, an example of 2d interpolation of an image is carried out using a combination
of PyLops operators (pylops.Restriction and
pylops.Laplacian) and the pylops.optimization module.
Mathematically speaking, if we want to interpolate a signal using the theory of inverse problems, we can define the following forward problem:
where the restriction operator \(\mathbf{R}\) selects \(M\) elements from the regularly sampled signal \(\mathbf{x}\) at random locations. The input and output signals are:
with \(M \gg N\).
import matplotlib.pyplot as plt
import numpy as np
import pylops
plt.close("all")
np.random.seed(0)
To start we import a 2d image and define our restriction operator to irregularly and randomly sample the image for 30% of the entire grid
im = np.load("../testdata/python.npy")[:, :, 0]
Nz, Nx = im.shape
N = Nz * Nx
# Subsample signal
perc_subsampling = 0.2
Nsub2d = int(np.round(N * perc_subsampling))
iava = np.sort(np.random.permutation(np.arange(N))[:Nsub2d])
# Create operators and data
Rop = pylops.Restriction(N, iava, dtype="float64")
D2op = pylops.Laplacian((Nz, Nx), weights=(1, 1), dtype="float64")
x = im.ravel()
y = Rop * x
y1 = Rop.mask(x)
We will now use two different routines from our optimization toolbox to estimate our original image in the regular grid.
xcg_reg_lop = pylops.optimization.leastsquares.NormalEquationsInversion(
Rop, [D2op], y, epsRs=[np.sqrt(0.1)], returninfo=False, **dict(maxiter=200)
)
# Invert for interpolated signal, lsqrt
(
xlsqr_reg_lop,
istop,
itn,
r1norm,
r2norm,
) = pylops.optimization.leastsquares.RegularizedInversion(
Rop,
[D2op],
y,
epsRs=[np.sqrt(0.1)],
returninfo=True,
**dict(damp=0, iter_lim=200, show=0)
)
# Reshape estimated images
im_sampled = y1.reshape((Nz, Nx))
im_rec_lap_cg = xcg_reg_lop.reshape((Nz, Nx))
im_rec_lap_lsqr = xlsqr_reg_lop.reshape((Nz, Nx))
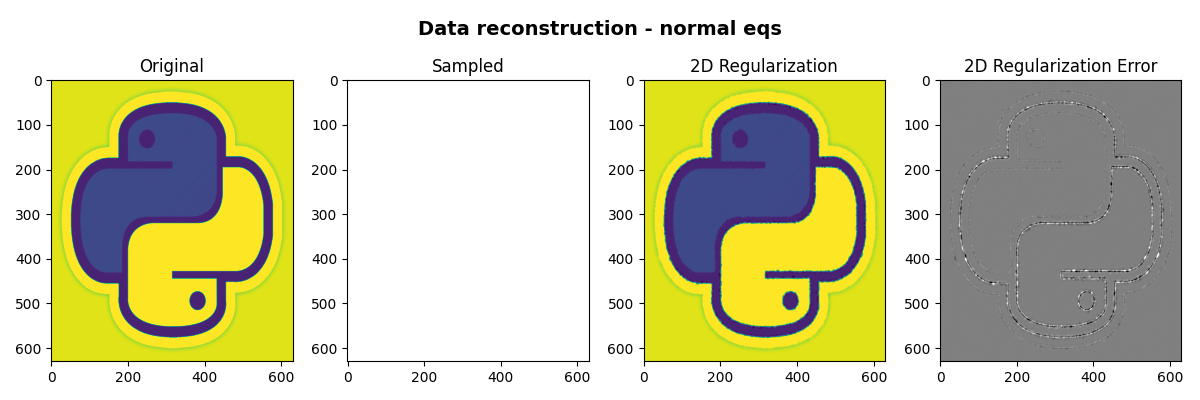
Finally we visualize the original image, the reconstructed images and their error
fig, axs = plt.subplots(1, 4, figsize=(12, 4))
fig.suptitle("Data reconstruction - normal eqs", fontsize=14, fontweight="bold", y=0.95)
axs[0].imshow(im, cmap="viridis", vmin=0, vmax=250)
axs[0].axis("tight")
axs[0].set_title("Original")
axs[1].imshow(im_sampled, cmap="viridis", vmin=0, vmax=250)
axs[1].axis("tight")
axs[1].set_title("Sampled")
axs[2].imshow(im_rec_lap_cg, cmap="viridis", vmin=0, vmax=250)
axs[2].axis("tight")
axs[2].set_title("2D Regularization")
axs[3].imshow(im - im_rec_lap_cg, cmap="gray", vmin=-80, vmax=80)
axs[3].axis("tight")
axs[3].set_title("2D Regularization Error")
plt.tight_layout()
plt.subplots_adjust(top=0.8)
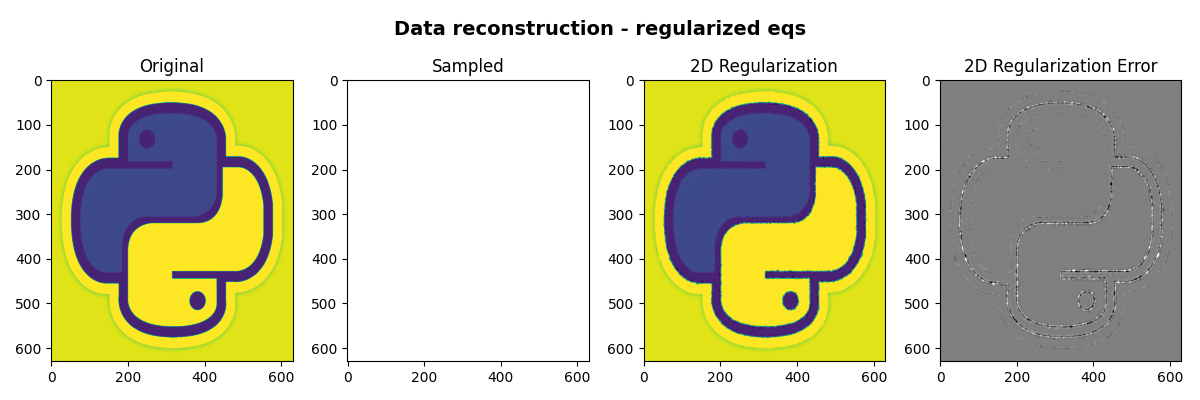
fig, axs = plt.subplots(1, 4, figsize=(12, 4))
fig.suptitle(
"Data reconstruction - regularized eqs", fontsize=14, fontweight="bold", y=0.95
)
axs[0].imshow(im, cmap="viridis", vmin=0, vmax=250)
axs[0].axis("tight")
axs[0].set_title("Original")
axs[1].imshow(im_sampled, cmap="viridis", vmin=0, vmax=250)
axs[1].axis("tight")
axs[1].set_title("Sampled")
axs[2].imshow(im_rec_lap_lsqr, cmap="viridis", vmin=0, vmax=250)
axs[2].axis("tight")
axs[2].set_title("2D Regularization")
axs[3].imshow(im - im_rec_lap_lsqr, cmap="gray", vmin=-80, vmax=80)
axs[3].axis("tight")
axs[3].set_title("2D Regularization Error")
plt.tight_layout()
plt.subplots_adjust(top=0.8)
Total running time of the script: ( 0 minutes 26.242 seconds)